The same algorithm used to find tunes in music retrieval systems has been successfully applied in identifying the signature whistles of dolphins, affording a new time-saving device for research into the world of dolphin communication.
Bottlenose dolphins, in particular, recognize each other by name: the sound of each animal’s “signature” whistle, which each dolphin develops at a young age. Bottlenose dolphins appear to show preference to the signature whistles of familiar individuals. Scientists have found that dolphins use the signature whistles to foster and maintain group cohesion.
Until now, the classification of individual dolphin whistles has typically been done by examining a spectrograph, which visually represents the spectrum of frequencies found in a sound. But the method is time-consuming, requires more data than might be necessary, and is subject to human error.
A study, published Oct. 23 in the journal PLOS ONE, describes a new method for identification that uses an algorithm based on what’s called the Parsons code, which has been used extensively in computerized retrieval of tunes from music databases.
I have a vested interest in Parsons code technology right now as one of my current machine learning projects relates to identifying individual music segments embedded in musical pieces, television shows, films, etc. This new research could point to a new path in identifying audio database content.
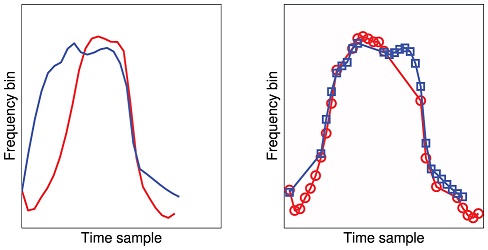
Instead of looking at the precise variation in frequency, the Parsons code only considers whether at each point in time the pitch goes up, down, or stays the same. The researchers examined 400 signature whistles from 20 different dolphins. The new algorithm did well at assigning signature whistles to individual animals, helping scientists to classify the tested whistles quickly and efficiently, according to the study.
The Parsons code is a robust way to compare dolphins’ signature whistles because it is able to home in on the variation in frequency that actually matters. It discards the information that isn’t useful for the analysis,” said lead author Arik Kershenbaum, a postdoctoral fellow at the National Institute for Mathematical and Biological Synthesis.




Speak Your Mind